MERFISH is a massively multiplexed single-molecule imaging technology for spatially resolved transcriptomics capable of simultaneously measuring the copy number and spatial distribution of hundreds to tens of thousands of RNA species in individual cells.
Accelerate to discover
Related topics
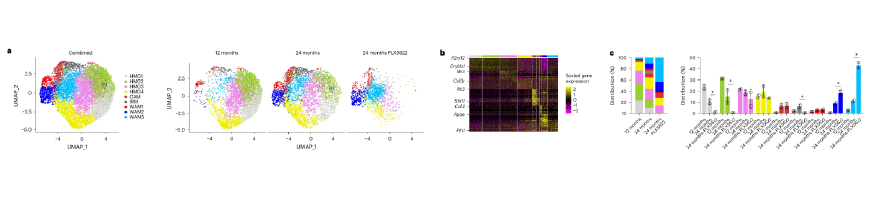
Use of Vizgen’s MERSCOPE Platform to Uncover New Insights about Aging
MERFISH and data analysis: Optic nerves were cut as 10-µm-thick transverse or longitudinal sections and collected onto glass slides supplied by Vizgen. To ensure proper adhesion, the glass slides were coated with a solution of 0.1 mg ml−1 poly-d-lysine bromide (Sigma-Aldrich, P7886), left to incubate at room temperature for 1 h and dried before starting the sectioning. The gene panel used in this study included 496 protein-coding genes, complemented by 54 blank probes (Supplementary Table 3). Within this gene panel, a diverse array of transcripts comprising recognized markers for glial and immune cell types was included based on previously published literature. Sections were postfixed with 4% PFA in PBS for 15 min at room temperature, washed three times with PBS and then placed in the autofluorescence bleacher (Vizgen) for 3 h52. Afterward, the sections were permeabilized in 70% ethanol at 4 °C overnight. For hybridization with the gene panel, samples were washed with a sample prep wash buffer (Vizgen) and then incubated in a formamide hybridization buffer at 37 °C for 30 min before adding 50 μl of the gene panel mix on top of the tissue. Sections were hybridized at 37 °C for 36–48 h, washed and embedded into a polyacrylamide gel according to the manufacturer’s instructions. Samples were cleared with Clearing Premix containing proteinase K, washed and stained with DAPI and polyT reagent for 15 min at room temperature. The samples were then washed, and the appropriate hybridization and imaging buffers were loaded onto the MERSCOPE system (Vizgen). A low-resolution mosaic was acquired using a ×10 objective, and regions of interest were selected for high-resolution imaging with a ×60 lens. For high-resolution imaging, the focus was locked to the fiducial fluorescent beads on the coverslip. Seven 1.5-μm-thick z planes were imaged for each field of view. Raw images were decoded to RNA spots with spatial coordinates and gene IDs using Merlin software (Vizgen) on the MERSCOPE instrument.
Related technologies: Spatial Transcriptomics



